The Blueprint of Living Things
How innovation at Los Alamos helped launch the Human Genome Project and the genomics revolution.
- Rebecca McDonald, Science Writer

In March 1986, a group of scientists (including several from Los Alamos) gathered in Santa Fe, New Mexico, to discuss the possibility of sequencing the entire human genome. Until then, while exciting and potentially valuable, the endeavor had not been considered feasible. Unraveling the blueprint of human beings could improve understanding of genetic disease and cancer, leading to advances in health care and prevention. People worried, however, that sequencing the genome would cost too much, that the technology was not readily available, and that it would require a level of international collaboration beyond anything scientists had done before.
The Santa Fe meeting was pivotal. It generated exciting ideas and momentum about how to address the technology challenges, and the scientists—including Los Alamos’s Scott Cram, Larry Deaven, and Robert Moyzis, who were awarded the Los Alamos Medal in 2017 for this work—helped the program leaders develop a plan forward. In 1987, the Department of Energy (DOE) began moving forward with a pilot Human Genome Initiative. Not everyone in the scientific community was excited, in fact, dozens wrote letters in opposition citing concerns about the “big science” nature of the project and the fact that a large percentage of the genome was likely “junk DNA.” Despite opposition within the National Institutes of Health (NIH), in 1988, the agency partnered with the DOE in the congressionally funded official Human Genome Project (HGP)—which today is regarded as one of the greatest achievements in science. The HGP has led to innumerable advances for human health and for understanding our relationship with the living world.
Why DOE?
Although it seemed unusual to many that the DOE was taking the lead to sequence the human genome, Los Alamos scientists didn’t think it was unusual at all: they had been laying the groundwork for decades.
We started the Human Genome Project without knowing what the technology would be like when we finished it
Beginning with the Manhattan Project, the DOE’s predecessor, the Atomic Energy Commission, was charged with conducting research on the health effects of radiation. By then, radiation had been linked to cancer, but scientists did not understand the underlying mechanisms or if these cancers would be heritable to the next generation. The Commission wanted to understand how radiation could impact its employees, the military, and anyone or anything exposed to it.
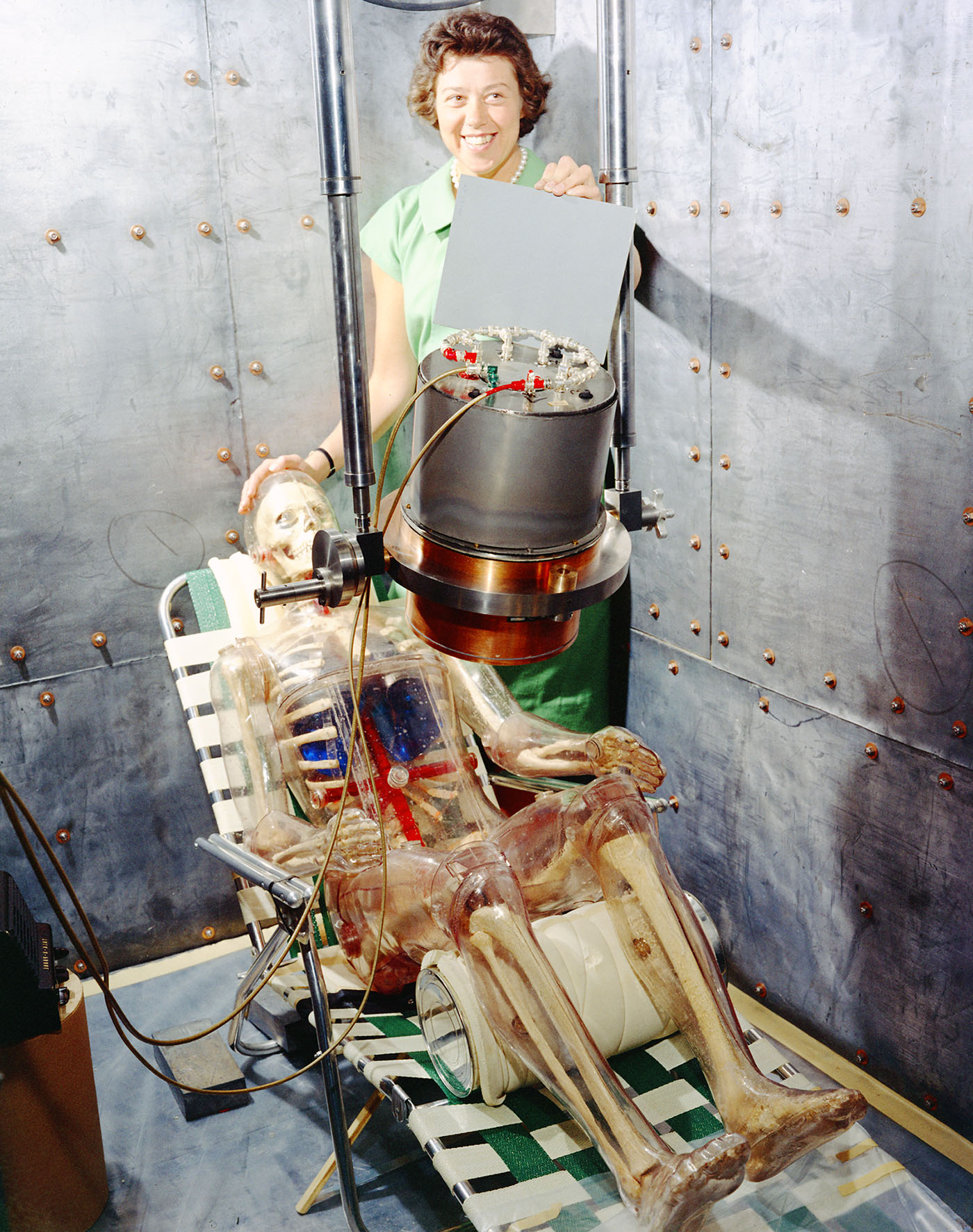
During the 1940s and 1950s, scientists at Los Alamos investigated this idea of “health physics.” In one project, to explore the effects of whole-body radiation exposure, Los Alamos chemist Wright Langham built “Plastic Man,” a life-sized dummy made to hold real bone and tissue samples. As biology advanced and scientists learned more about the molecular make-up of organisms (such as the 1953 discovery of DNA’s structure), it became logical to change focus from animal tissues to the cells and molecules within them. Perhaps understanding DNA could elucidate how it might be impacted by radiation.
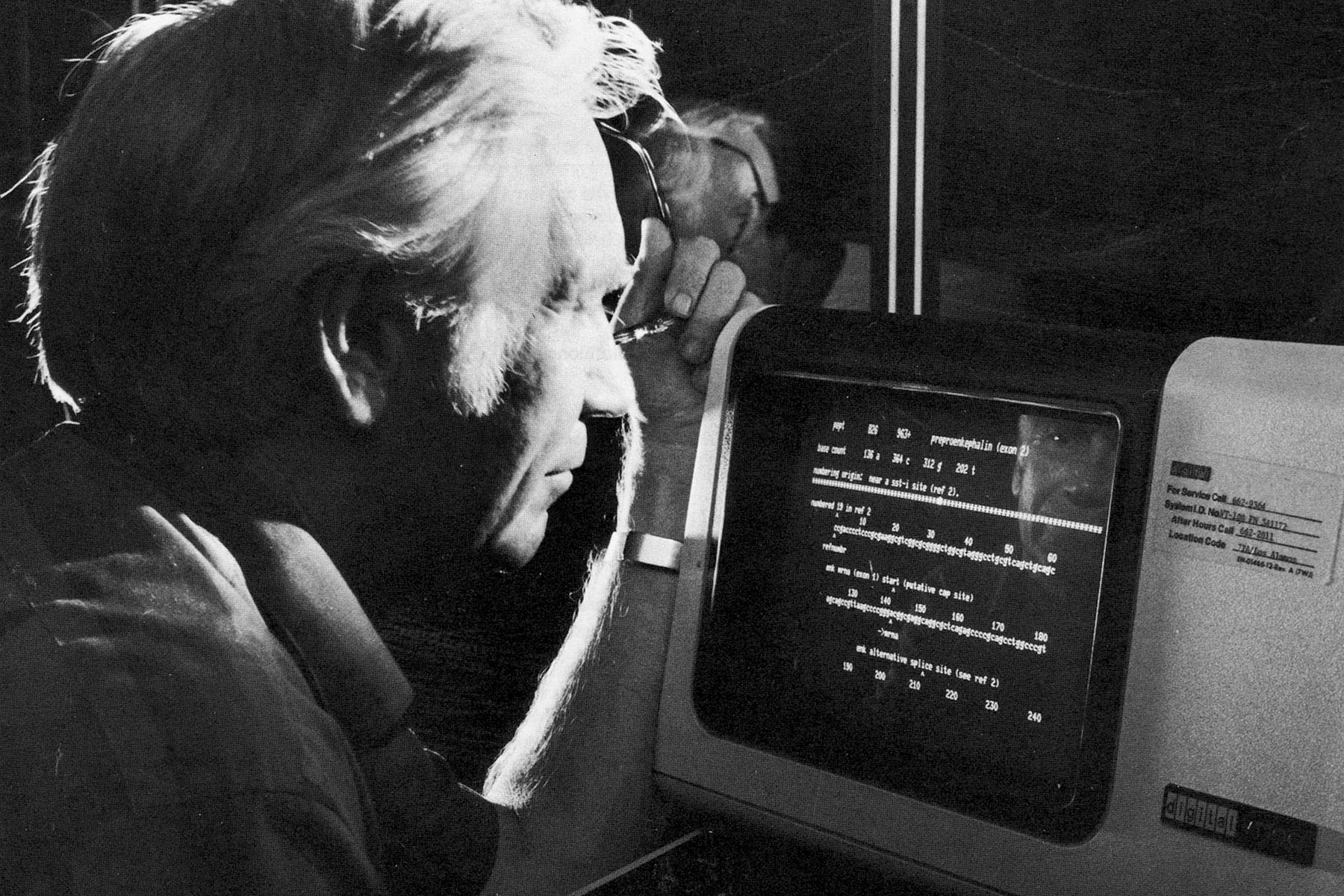
Over the next few decades, several distinct projects happening in parallel at Los Alamos helped lay the foundation for the technologies used in the HGP for sequencing and analysis. In the 1960s, scientists figured out how to read DNA by deciphering its sequences. By the 1970s, sequencing was becoming widespread in the scientific community and a handful of Los Alamos physicists—who had turned their attention to biology and computation—realized that the accumulating sequence data would need a repository. Physicist Walter Goad and his team established the Los Alamos Sequence Database in 1979, populating it with published sequences and developing the computational framework to efficiently share this collection with the greater scientific community for comparative analysis. In 1982, the database was renamed GenBank, and it was housed at Los Alamos until 1992 when it was moved to the NIH. GenBank now holds more than 4.7 billion sequences for more than half a million species.
In another Los Alamos group, in the 1960s, scientist Mack Fulwyler made a device to count, sort, and identify individual cells. This device led to the development of modern-day flow cytometers and cell sorters, which are now ubiquitous throughout medical and research laboratories worldwide. In 1982, Los Alamos began the NIH-funded National Flow Cytometry and Sorting Research Resource through which scientists developed new types of flow cytometers and research techniques. In addition, Los Alamos joined with Lawrence Livermore National Laboratory for the DOE National Laboratory Gene Library Project, in which they used flow cytometry to create copies of human chromosomes for distribution to researchers. This capability was critical in the early days of the HGP because it ensured that enough copies of a specific chromosome’s DNA would be available for sequencing. During the HGP, Los Alamos played a key role in providing chromosome-specific DNA libraries to partner institutions, while focusing in-house efforts on the sequencing of chromosome 16 and parts of chromosome 5 (each participating institution was assigned different chromosomes).
Beyond its role in the HGP, Los Alamos was a founding partner in the DOE’s Joint Genome Institute (JGI) in 1997, and Lab scientists continued to hone sequencing protocols, curate databases, and develop the new field of bioinformatics to computationally turn the increasing amount of data into actionable information.

Next generation genomics
When the HGP officially launched, DNA sequencing was a tedious, labor-intensive process that cost a dollar or more per base pair, putting the price tag of the entire project to over three billion dollars.
“We started the HGP without knowing what the technology would be like when we finished it,” says biologist and Laboratory Fellow Babetta Marrone. “We needed the project to revolutionize the tech.”
A technology revolution is precisely what happened. The HGP was expected to take 15 years, but after several twists and a lot of drama, the genome was delivered two years early, in 2003, along with a disruptive new technique called shotgun sequencing that transformed the future of genome science completely. Every year that followed saw advances in sequencing machines and techniques, making it easier and exponentially less expensive to sequence genomes. Today, the cost per base pair is approximately one tenth of a cent.

Throughout this revolution, Los Alamos scientists, the JGI, and several other institutions began applying the new sequencing methods and technologies to a variety of microorganisms and animals to more thoroughly investigate their evolutionary relationships, how they function, and how they interact. In addition, building on the idea of GenBank, Lab computational biologists compiled species-specific databases, the first one being a Human Immunodeficiency Virus (HIV) database created in 1986 by Lab scientist Gerald Myers in response to the AIDS pandemic. Recognizing HIV’s extraordinarily rapid rate of evolution, the data helped researchers track the virus’s evolution and diversity. Myers’s database enabled the comparative analysis of pathogen genomes, leading to strategies for vaccine and therapeutics development, and informed epidemiological tracking of viruses through populations.
Today, genomics helps scientists investigate microorganisms, plants, animals, and humans—their evolutionary relationships, how they function, and how they interact
In its earliest days, the HIV database was provided to researchers annually as a hardcopy book. Computational biologist and Lab Fellow Bette Korber, who led the HIV database for 25 years after Myers retired, remembers visiting labs where the book was actually chained to people’s desks to keep others from walking off with it. Today, the HIV database is available online with close to a million HIV sequences, and the Lab hosts additional databases for Human Papilloma Virus, influenza, Hepatitis C, ebolavirus, and even SARS-CoV-2, the virus that causes COVID-19.
Omniscient ‘omics
Because of the technological advances spurred by the HGP, scientists can now routinely evaluate new genomes (genomics), RNA and gene regulation (transcriptomics), chemical modifications to genes that can influence their regulation (epigenomics), and identify encoded proteins (proteomics) and chemical signals (metabolomics). These technologies, used independently, or together (called multi-omics), allow Los Alamos scientists to explore human disease and normal function, host-pathogen interactions within an organism, and the spread of pathogens through humans and animal populations.
“As SARS-CoV-2 was just beginning its global expansion, we rapidly adapted computational tools we had first developed to study HIV sequence evolution, and this enabled us to discover, in 2020, that SARS-CoV-2 was evolving to be more infectious,” says Korber. Throughout the COVID-19 pandemic, Los Alamos scientists helped track variants, supporting discoveries in near-real-time regarding how specific mutations impacted vaccine efficacy, transmission, and drug and antibody resistance.
Los Alamos scientists have also helped pioneer the new field of microbiome science, which is the study of the assemblage of microbes that live within an environment such as a human gut or skin, an algae pond, or a corn field. The ability to elucidate threats in these types of complex environments profoundly enhances the nation’s security through understanding the conditions and organisms that promote healthy plants and animals compared to those that cause disease.
“We are employing traditional bioinformatics and new tools driven by AI, to analyze in real time, large volumes of sequencing data,” explains computational biologist and Lab Fellow Patrick Chain. “For instance, using data from an environment like wastewater we can try to anticipate novel and emerging pathogens, predict their potential for spread, their evolution, and what diagnostics and countermeasures can be deployed to mitigate their impact.”
It is unlikely that the scientists irradiating Plastic Man in the 1950s anticipated that their work would begin a trajectory towards tracking biothreats through global populations and their sewers, but it is an undeniable example of how basic scientific research feeds an unlimited future of discovery.
People Also Ask:
- How many genes are contained in the human genome? Humans have approximately 20,000 protein-coding genes.
- What is the Human Genome Project? The Human Genome project was a large, international effort founded in 1990 by the US Department of Energy and the National Institutes of Health to map and sequence all the genes in human DNA: the human genome.








